tfcb_2020
Lecture 4: Reproducible Research, Markdown, Git and GitHub
Trevor Bedford (@trvrb, bedford.io)
Learning objectives
- Identify minimum requirements for a reproducible computational project
- Apply basic tasks using Git and GitHub to track versions of files
- Write formatted text files using Markdown
Class materials
Reminders
- Recommended reading is available at the end of each section
- Your second homework assignment is available today, and due Tuesday, October 20 at 1pm. This homework covers project organization and Git/GitHub (lectures 4) as well as tidy data (lecture 5). Contact Kate (khertwec at fredhutch.org) with any questions or concerns.
Reproducible science
Motivation
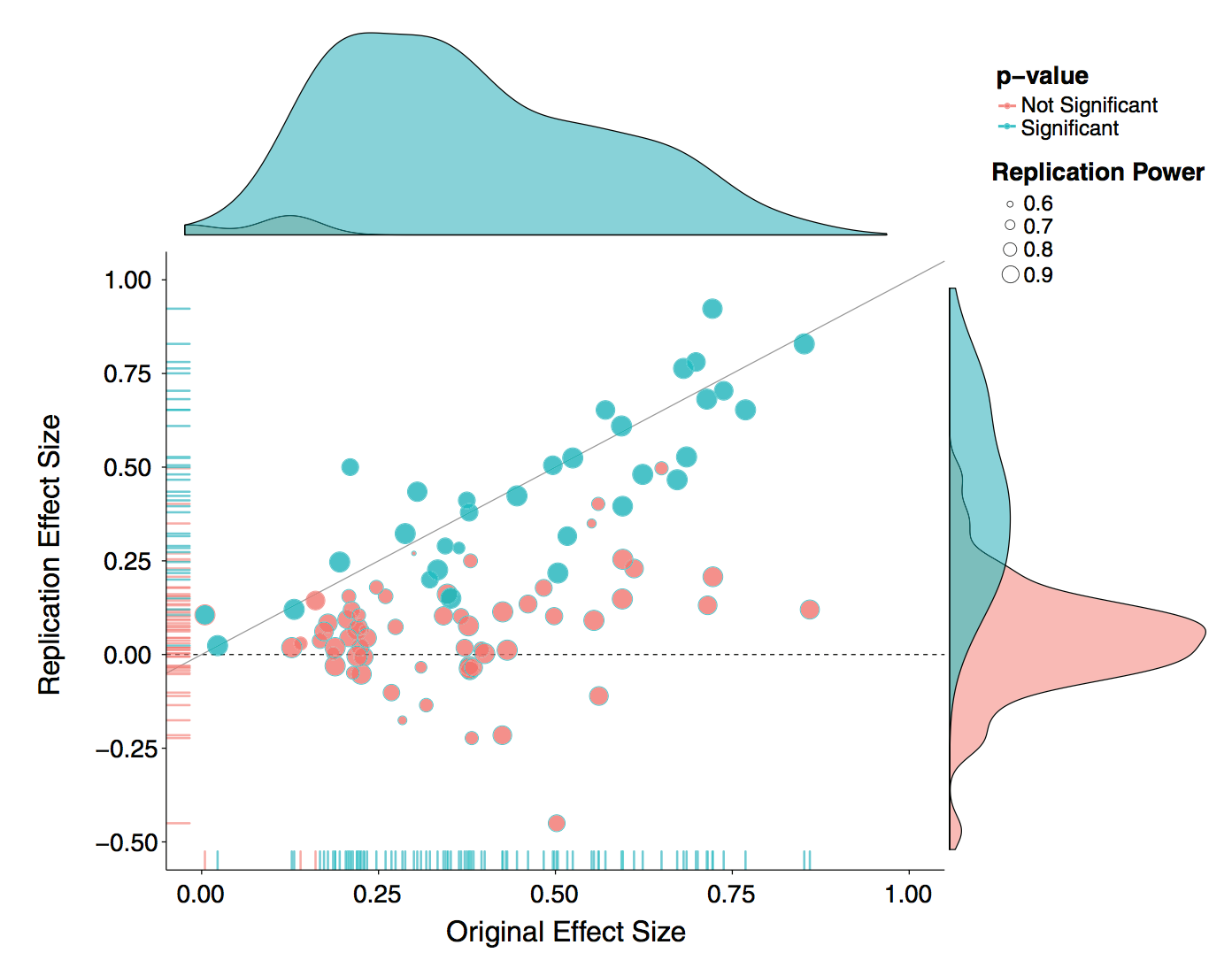
There is a lot of interest and discussion of the reproducibility “crisis”. In one example, “Estimating the reproducibility of psychological science” (Open Science Collaboration, Science 2015), authors attempt to replicate 100 studies in psychology and find that only 36 of the studies had statistically significant results.
The Center for Open Science has also embarked on a Reproducibility Project for Cancer Biology, with results being reported in an ongoing fashion.
There are a lot of factors at play here, including “p hacking” lead by the “garden of forking paths” and selective publication of significant results. I would call this a crisis of replication and have this as a separate concept from reproducibility.
But even reproducibility is also difficult to achieve. In “An empirical analysis of journal policy effectiveness for computational reproducibility” (Stodden et al, PNAS 2018), Stodden, Seiler and Ma:
Evaluate the effectiveness of journal policy that requires the data and code necessary for reproducibility be made available postpublication by the authors upon request. We assess the effectiveness of such a policy by (i) requesting data and code from authors and (ii) attempting replication of the published findings. We chose a random sample of 204 scientific papers published in the journal Science after the implementation of their policy in February 2011. We found that we were able to obtain artifacts from 44% of our sample and were able to reproduce the findings for 26%.
They get responses like:
“When you approach a PI for the source codes and raw data, you better explain who you are, whom you work for, why you need the data and what you are going to do with it.”
“I have to say that this is a very unusual request without any explanation! Please ask your supervisor to send me an email with a detailed, and I mean detailed, explanation.”
Tables in paper are very informative.
At the very least, it should be possible to take the raw data that forms the the basis of a paper and run the same analysis that the author used and confirm that it generates the same results. This is my bar for reproducibility.
Reproducible science guidelines
My number one suggestion for reproducible research is to have:
One paper = One GitHub repo
Put both data and code into this repository. This should be all someone needs to reproduce your results.
Digression to demo GitHub.
This has a few added benefits:
-
Versioning data and code through GitHub allows you to collaborate with colleagues on code. It’s extremely difficult to work on the same computational project otherwise. Even Dropbox is messy.
-
You’re always working with future you. Having a single clean repo with documented readme makes it possible to come back to a project years later and actually get something done.
-
Other people can build off your work and we make science a better place.
I have a couple examples to look at here:
- My 2015 paper on global migration of influenza viruses: github.com/blab/global-migration
- Sidney Bell’s 2018 paper on dengue evolutionary dynamics: github.com/blab/dengue-antigenic-dynamics
Some things to notice:
- There is a top-level
README.md(written in Markdown… more to come) file that describes what the thing is, provides authors, provides a citation and link to the paper and gives an overview of project organization. - There is a top-level
data/directory. - Each directory should have a
README.mdfile describing its contents. - The readme files should specify commands to run, or point to the make file that does.
- Versioned Jupyter notebooks (example) are viewable in the browser, as are
.tsvfiles (example). - Relative links throughout.
- Include language-specific dependency install file, Python uses
requirements.txtas standard. - Figures are embedded directly in the readme files via Markdown (example).
More sophisticated examples will use a workflow manager like Snakemake to automate builds. For example:
- Alistair Russell and Jesse Bloom’s recent work on single-cell sequencing of influenza: github.com/jbloomlab/IFNsorted_flu_single_cell/
- John Huddelston’s influenza forecasting work: github.com/blab/flu-forecasting (this also provides a Conda environment for better reproducibility)
With GitHub as lingua franca for reproducible research, there are now services built on top of this model. For example:
- Zenodo allows you to mint DOIs from GitHub releases.
- In github.com/cboettig/noise-phenomena, Carl Boettiger provides
.Rmdof code, but also uses Binder to launch an interactive RStudio session. Binder is described in more detail here. - We’ve built Nextstrain to look for results files in public GitHub repos to provide interactive figures. For example nextstrain.org/community/blab/zika-colombia provides an interactive visualization directly from files hosted on GitHub at github.com/blab/zika-colombia/.
Project communication
For me, as PI, I enforce a further rule:
One paper = One GitHub repo = One Slack channel
It’s much easier if all project communication goes in one place.
Further reading
Some suggested readings on reproducible research include:
Markdown

-
What is Markdown? (Look at this lecture!)
-
Invented by John Gruber to provide markup that is intuitive and aesthetically pleasing when viewed as plain text. It’s similar to how you would style a plain text email.
-
Check out this Markdown cheatsheet. Also, GitHub’s Markdown style guide
-
Markdown is well supported by GitHub. Because Markdown is plain text it versions well. Word docs version horribly.
-
I strongly support its separation of content and styling.
-
Some further examples of Markdown:
- Practical I wrote on using BEAST
- Benchtop protocol for Zika sequencing written by Alli Black
- bedford.io is written completely in Markdown (example)
Git and GitHub

-
Why use version control?
-
Introduction to Git and distributed version control: Basics from Alice Bartlett with Git for Humans
-
Introduction to GitHub
-
GitHub Desktop vs command line
- Demonstration:
- Make a new project on GitHub
- Make a linear series of commits in GitHub Desktop
- Cover same actions on the command line:
git status,git add,git commit,git diff - Safely exploring history and rolling back if necessary:
git log,git checkout - Remotes and push and pull from GitHub:
git pull,git push - Look at files under the hood
- Other details:
- The importance of
.gitignore - The importance of good commit messages, relevant xkcd
- Breaking up work and staging / committing different pieces separately
- Merge conflicts (don’t panic)
- The importance of